Summary:
Bacteria found in freshwater lakes can originate in soil and adapt through different evolutionary routes, as shown in a study published in Nature Communications. Researchers at the University of Zurich (UZH) examined the genomes of Limnocylindria bacteria in Lake Zurich, identifying how these microbes transitioned from terrestrial to aquatic environments.
The analysis revealed two contrasting strategies. One lineage increased its genome size by acquiring genes that enable survival in water, including traits linked to movement. Another lineage took the opposite approach, reducing its genome and eliminating functions no longer needed in freshwater conditions. This simplified group requires fewer resources and has become widespread in the lake.
The study also identifies genetic processes behind these changes, including mutation-driven genome restructuring combined with selective pressure for efficient use of carbon and nitrogen in proteins. Both gene gain and gene loss can support movement across ecosystems, providing a clearer understanding of how bacteria establish themselves in new environments.

— Press Release —
How soil microbes adapt to life in lakes
Bacteria are tiny – and incredibly old: they were among the first life forms to emerge on our planet around four billion years ago. Since then, they have “infected the entire Earth,” says Adrian-Stefan Andrei, head of a research group at the Department of Plant and Microbial Biology at the University of Zurich’s Limnological Station. “They swim in the oceans, are found deep in the soil as well as in and on other living organisms – and some of them even float high up in the atmosphere.”
Traces in the genome
But how do bacteria manage to conquer new environments? “This question remains largely unanswered, partly because very few microbes leave fossil traces,” says Andrei. He and his team have now used bioinformatic methods to get closer to answering this question. “We have examined the traces that evolution leaves in the genomes of living organisms,” he says.
Andrei’s team compared the genomes of representatives of a specific class of bacteria called Limnocylindria, finding that these bacteria originally lived in the soil. In a recently published paper, the researchers report that even today many bacteria of the Limnocylindria class populate the dark soil deep beneath our feet. However, two subgroups can also be found in freshwater lakes such as Lake Zurich.
Additional genes for special traits
Members of one subgroup – with the unwieldy name CSP1-4 – have a larger genome compared to their soil-dwelling relatives. Their genetic material also contains genes that enable them to produce so-called flagella, which are fine filaments on the surface that rotate like propellers and enable the bacteria to move through the water. The genome of CSP1-4 bacteria tells a rather unsurprising story, says Andrei. The additional genes confer special properties on the microbes, enabling them to survive in their new environment.
Simplifiers have already played their hand
The genetic analysis of the other subgroup (Limnocylindraceae), however, yielded unexpected results: the genome of these freshwater bacteria is only half the size of that of their soil-dwelling counterparts. This group – which Andrei refers to as the simplifiers – underwent the opposite evolution when they moved from land to water: instead of acquiring new genes, these microbes shed a significant portion of their genome. “They got rid of their unnecessary genetic luggage – and travelled lighter as a result,” says Andrei.
Today, these simplifiers are abundant in Lake Zurich. “They require few resources. That is the secret to their current, very great ecological success,” says Andrei. However, unlike the CSP1-4 group, they are now only found in freshwater. The simplifiers seem to already have played their hand. Due to their reduced genome, they are unlikely to ever be able to leave their freshwater home again. So, according to Andrei, there is no blanket answer as to whether it is better for bacteria to enlarge or reduce their genome before colonizing a new environment.
Journal Reference:
Serra Moncadas, L., Shakurova, A., Hofer, C. et al., ‘Deep-branching Chloroflexota lineages illuminate the eco-evolutionary foundation of cross-ecosystem colonization’, Nature Communications (2026). DOI: 10.1038/s41467-026-71228-y
Article Source:
Press Release/Material by University of Zurich (UZH)
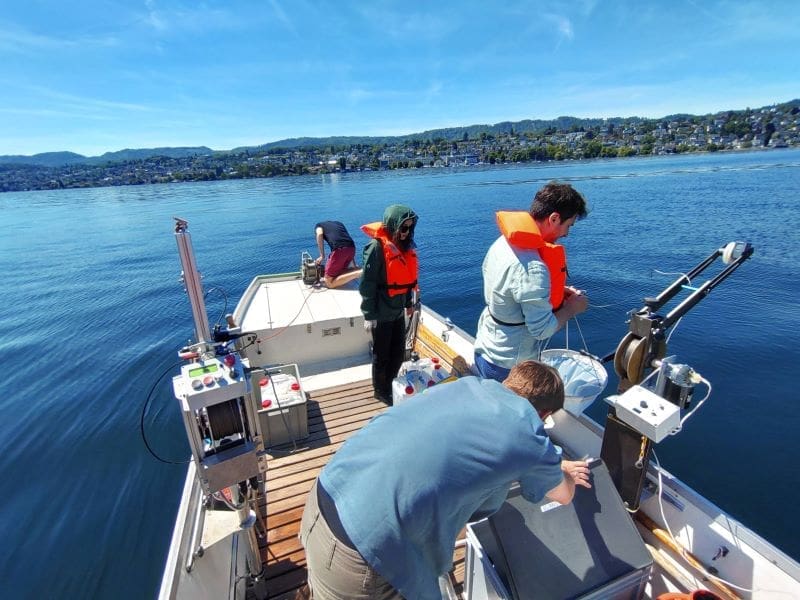
Featured image: UZH researchers from the Limnological Station conducting microbial monitoring on Lake Zurich during a field campaign: Water samples are collected using specialized equipment for downstream ecological and molecular analyses. Credit: Gianna Dirren-Pitsch | UZH